 P.S: I incubated these membranes with primary antibody A, read it, then primary antibody B, read it, then lastly with housekeeping gene. 
Thus, I am now very confused of which program is more reliable. And my PI does not agree that ImageJ makes more sense than Odyssey. This time the numbers are much similar to each other, but I still don't understand how the area and percent work and what they really mean. Then I tried using ImageJ to re-do all measurements. If even the housekeeping gene readings are so different, how can we expect the soluble protein readings to be accurate? (this is a question) After transforming a source image into an 8-bit greyscale image and inverting it in ImageJ/Fiji, the rectangle tool and ROI manager were used to define a series of regions of interest (ROIs), covering each dot with a single ROI of the same size, and the integrated density of each ROI was. using different quantification softwares and techniques, and then comparing the. Protein Quantification with Fiji and Microsoft Excel. Same tools can be used for DNA gel images, thin layer chromatography, Southern blot. This guide summarizes the basic principles of normalization and explains. This demonstrates how to create a figure to display western blot results. Recent changes, notably updates to the Journal of Biological Chemistrys submission guidelines (1), necessitate a closer look at traditionally accepted normalization practices. For example, I ran 40ug of proteins for all samples, but I would get an average intensity reading of 80 on one membrane and 120 on another for the housekeeping gene. From google search, there are three methods to do background subtraction in analyzing western blots with ImageJ. Normalization is a critical part of attaining reproducible data from quantitative Western blots. However, no matter how similar the procedures were, there was still some huge difference in band expression. We are trying to compare between 2 sets of 8 samples, so I ran 2 western blots simultaneously all the time. Not sure if many people are familiar with the Odyssey software, but we use it to scan our membrane and to quantify the bands. I am currently using IR-dye secondary antibodies purchased from LI-COR to detect some soluble proteins. Copy the results into an excel file and analyse as necessary.Hi all, this is my very first project and I hadn't have any experience at all.1.49K subscribers Subscribe Subscribed 152 Share 35K views 2 years ago Data Analysis Subscribe. Be sure to count the peaks on the plot as to determine which peak corresponds to the peak of interest. Quantification of the protein bands was done using Fiji software (ImageJ, NIH), free software with the integrated plug-in for Western blot quantification (Gassmann et al., 2009). HOW TO QUANTIFY WESTERN BLOT BANDS using ImageJ Area Under The Peak Method Adwoa Biotech. Make sure that the image is in 8-bit mode: go to Image>Type>8-bit. 
Go to the results window and the areas are calculated in numerical order. Analyzing gels and western blots with ImageJ 1. HOW TO QUANTIFY WESTERN BLOT BANDS using ImageJ Area Under The Peak Method Adwoa Biotech.Click on selected peaks (all peaks which are above the baseline) in order. You will use this program to determine the relative quantities of protein in each experimental sample. 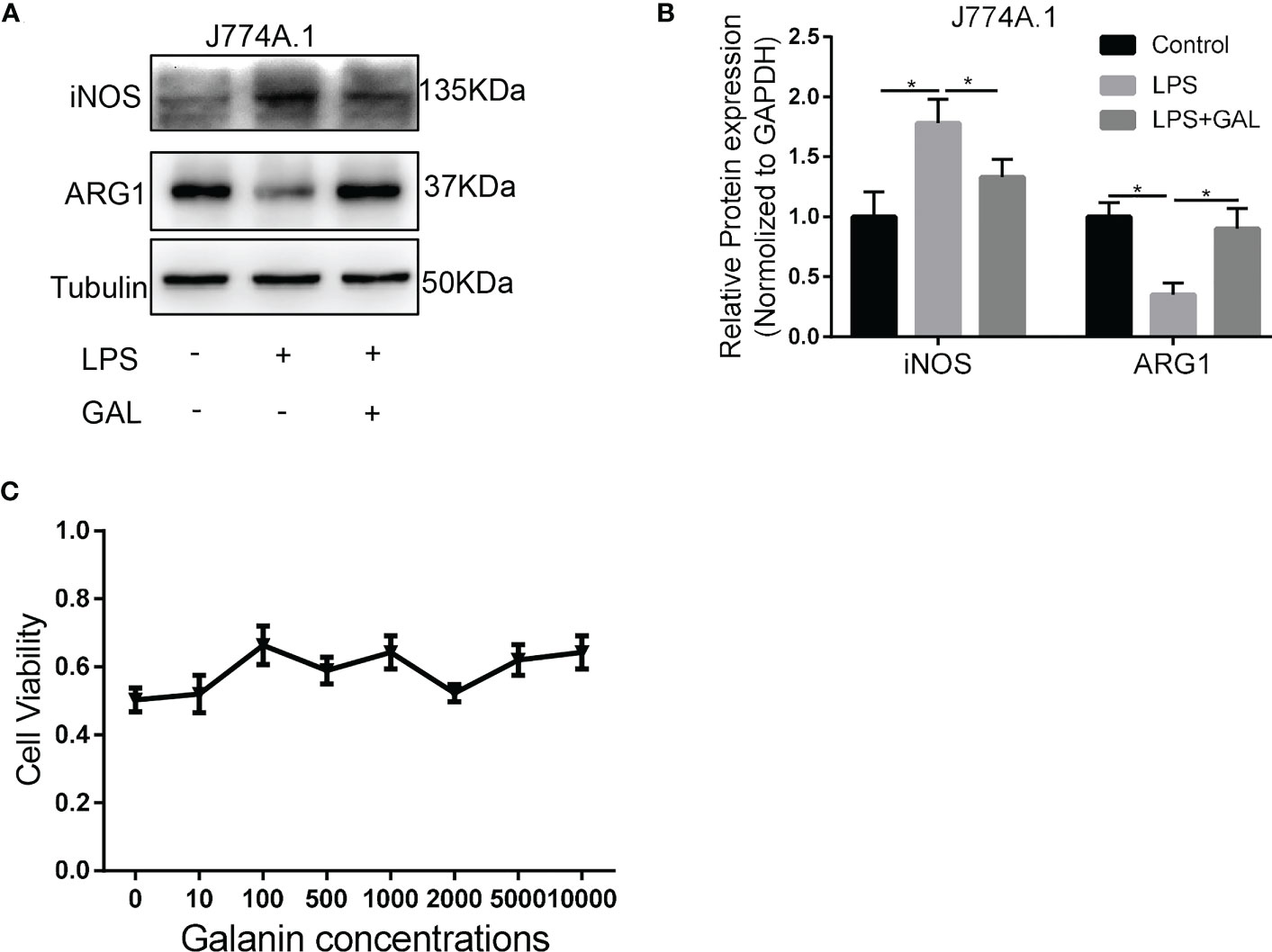
To quantify my western blot bands: I need to place a box around each band Then go to Anayze>Gels>Plot lanes From the generated plots I mark the bell shaped curve with the line tool Use the wand and get a quantification value from the intensity of my band. Western Blot Image Analysis: Lane and Band Tools.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
